INTRODUCTION
The use of pesticides in Brazil is widespread and expanding. According to an Ibama report, sales of formulated products in 2009 reached 306000 tons. In 2022, after a gap of 13 years, sales of 800000 tons were recorded, which represents an increase of 261.43%. Among the various chemical compounds under analysis, glyphosate and its salts have the highest sales volume, with a total of 266000 tons sold1. Glyphosate, an aminophosphate compound, is used as a broad-spectrum herbicide and was first commercially synthesized in 1970. By 2010, the compound had been released for use in more than 130 countries. Glyphosate acts on the enzyme 5-enolpyruvylshikimate-3-phosphate synthase (EPSPS), inhibiting its function. This enzyme is present in fungi, plants, and bacteria2. The International Agency for Research on Cancer (IARC) has re-assessed the carcinogenic potential of glyphosate and other pesticides, maintaining them in category 2A as potentially carcinogenic3. Thus, there is a clear need for research into the possible effects on human and animal health.
A review on the subject is warranted due to the wide use of glyphosate, which is listed as a potentially carcinogenic via IARC. Therefore, a review of the subject would be of paramount importance to help organize the data obtained on the epigenetic effects linked to the agrochemical. The aim of this narrative review was to summarize and evaluate the experimental results available in scientific databases that relate the exposure of cells and animal models to glyphosate with epigenetic and toxicogenomic effects.
In this narrative review all articles included dealt with glyphosate, or formulations based on glyphosate or its main metabolite, AMPA. The aim of this review is to highlight the epigenetic effects of glyphosate, a notable distinction being the integration of studies employing human cell populations or animal models. Only articles published between 2018 and 2024 were searched. None of the articles used for the review were reviews or letters to the editor. The articles used were in English. The guiding question of this review was: ‘What are the toxicogenomic and epigenetic effects of glyphosate in animal and cell culture models?’. The objective of this review is to select in vitro experiments with human cell cultures and/or animal models exposed to glyphosate that analyze epigenetic effects in these populations. This review was registered in PROSPERO (ID 571598), and the protocol was last edited on 22 July 2024. This narrative review followed the PRISMA guidelines4.
The searches were carried out in scientific databases: PubMed, Capes Journals, Virtual Health Library Brazil (BVS) and Lilacs via BVS from 2018 to 2024. The terms used for the search were: ‘Glyphosate’, ‘Toxicogenomics’ and ‘Epigenetics’. The terms were organized into two searches using the Boolean operator AND: ‘Glyphosate’ AND ‘Epigenetics’, and ‘Glyphosate’ AND ‘Toxicogenomics’.
First, articles on the topic of interest were selected by title and abstract screening, and any discrepancies or doubts were discussed with a third reviewer based on the inclusion/exclusion criteria. After abstract and title screening, the remaining were screened based on their full text. The reviewers extracted information regarding the experimental models (human cell lines or animal models), the specific glyphosate formulation used (pure glyphosate vs glyphosate-based herbicides, i.e. Roundup®), dosages, and routes of exposure. Synthesis of the data was performed using a functional thematic approach, where results were categorized into specific epigenetic markers: global methylation, DMRs, epigenetic enzyme activity, hormonal disruption, tumor-related gene expression, and general toxicogenomic profiles. Due to a lack of standardization in experimental models and applied doses across the 20 selected studies, a narrative synthesis was applied rather than performing a quantitative meta-analysis.
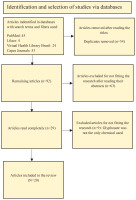
A total of 126 articles were found. Of this total, 34 were duplicates across databases. Of the remaining 92 that were not duplicates, 63 were removed after reading their abstracts. After exclusion, 29 remained. Of these 29, six were removed after reading full texts, as they did not fit the scope of the review. The main reason for excluding the articles was that they did not study glyphosate independently, making it impossible to isolate the effects caused exclusively by glyphosate. The articles excluded from the review moreover were not utilizing human cells and/or animal models, or had not yet conducted epigenetic analyses. A further reason for the exclusion is that many articles did not explicitly indicate in their titles and abstracts that they were review articles, which were subsequently excluded. Finally, a total of 20 articles were included in this review.
EPIGENETIC AND TOXICOGENOMIC EFFECTS OF GLYPHOSATE
Global methylation
In the selected articles, in relation to global methylation, the methylation status was not significant in the study by Larsen et al.5 in pig models. In contrast, three studies showed hypomethylation, these being Serra et al.6, Woźniak et al.7,8 and Duforestel et al.9 with the experimental models studied being roosters, peripheral blood mononucleated cells (PMBC) and MCF10A cell line, respectively (Table 1). In the study with pigs5, the route of exposure to glyphosate was feed, and no significant results were found in the variation of the percentage of 5-methyl cytosine (5mC) in the livers, kidneys and small intestines at any of the concentrations tested, which were 20 ppm and 200 ppm (0.000019 kg/L and 0.00019 kg/L, respectively) of the agrochemical5. However, a significant decrease in the percentage of 5mC was observed in roosters whose route of exposure was also feeding, but in sperm. The roosters were treated with 46.8 mg/kg body weight/day of Roundup® (commercial glyphosate-based solution)6, which is below the maximum daily concentration where adverse effects are not observable for birds (NOAEL)10.
Table 1
Comparison of global methylation results
| Authors Year | Experimental model | Concentration | Compound | Methylation status |
|---|---|---|---|---|
| Larsen et al.5 2022 | Pigs (liver, kidney, and small intestine) (n=26) | 20 ppm | Glyphosate | Not significant |
| Larsen et al.5 2022 | Pigs (liver, kidney, and small intestine) (n=26) | 200 ppm | Glyphosate | Not significant |
| Serra et at.6 2021 | Roosters (sperm) (n=5) | 46.8 mg/kg/day | Roundup® | Hypomethylation |
| Woźniak et al.7 2019 | PBMC (healthy volunteers, n=4) | 0.085, 1.7 and 17 mg/L | Glyphosate | Hypomethylation |
| Woźniak et al.8 2020 | PBMC (healthy volunteers, n=4) | 1.7 and 42.5 mg/L | AMPA | Hypomethylation |
| Duforestel et al.9 2019 | MCF10A | 10-11 M (cell concentration used = 2.2×10-6) | Glyphosate | Hypomethylation |
In human tissues, changes in the global methylation pattern can also be assessed during exposure to glyphosate. As in the case of the article by Woźniak et al.7, in which PBMCs exposed to 0.5, 10 and 100 μM (0.085, 1.7 and 17 mg/L, respectively) of glyphosate obtained a significant drop in the percentage of 5mC at all concentrations tested. In the same study, PMBC also suffered global hypomethylation when treated with aminomethylphosphonic acid (AMPA), the main secondary metabolite of glyphosate, at concentrations of 10 and 250 μM (1.7 and 42.5 mg/L, respectively).
Differentially methylated regions
Τhe studies by Kubsad et al.11, Beck D. et al.12 and Ben Maamar et al.13, presented analyses of DMRs in models treated with glyphosate. In the study by Kubsad et al.11 a transgenerational analysis was carried out, female Sprague Dawley rats were treated with 25 mg/kg body weight/day during days 8 to 14 of pregnancy, the F1 generation (exposed during the fetal period) generated F2, which in turn generated F3. Analysis of DMRs in sperm showed that the number of DMRs varied in each generation, and that DMRs did not overlap. Therefore, no region had the same methylation status in all generations.
Beck D. et al.12 carried out a transgenerational study along the same lines as the article in the previous paragraph; however, only individuals who had disease in specific tissues at F3 were analyzed. In the work by Ben Maamar et al.13, the transgenerational approach was maintained. In the article, tissues from F3 individuals treated with glyphosate who developed diseases were for analysis of DMRs (kidney disease, prostate disease, obesity and multiple pathologies), in the work the individuals had the same numbers of DMRs as the work by Beck et al.12. Furthermore, they arrive at the result that a large part of the DMRs measured are related to signaling genes, metabolism, transcription and receptors.
Enzymes related to epigenetic mechanisms
A number of identified studies (Smith et al.14, Larsen et al.5, Woźniak, et al.15; Duforestel et al.9 and Courant et al.16) evaluated different enzymes with functions related to epigenetic mechanisms (Table 2).
Table 2
Comparison of results related to enzymes of epigenetic mechanisms
| Authors Year | Experimental model | Concentration | Gene | Compound | Expression |
|---|---|---|---|---|---|
| Smith et al.14 2019 | Oryzias latipes (n=20–37) fish | 5 mg/L | Dnmt1 | Roundup® | Decrease |
| Smith et al.14 2019 | Oryzias latipes (larvae) | 0.5 mg/L of glyphosate, and 0.5 and 5 mg/L of Roundup® | TET1 and TET3 | Glyphosate and Roundup® | Increase |
| Smith et al.14 2019 | Oryzias latipes (larvae) | 0.5 and 5 mg/L | TET2 | Roundup® | Increase |
| Larsen et al.5 2022 | Pigs (kidney, liver, and small intestine) (n=26) | 20 and 200 ppm | Dnmt1, Dnmt2, Dnmt3A, Dnmt3B and TET3 | Glyphosate | Not significant |
| Woźniak et al.15 2021 | PBMC (healthy volunteers, n=4) | 0.5, 10, and 100 µM | Dnmt1 | Glyphosate | Increase |
| Woźniak et al.15 2021 | PBMC | 100 and 250 µM | Dnmt1 | AMPA | Increase |
| Woźniak et al.15 2021 | PBMC | 100 µM | Dnmt3A | Glyphosate | Increase |
| Woźniak et al.15 2021 | PBMC | 5, 10, and 100 µM of glyphosate, and 100 and 250 µM of AMPA | EHMT1 and EHMT2 | Glyphosate and AMPA | Not significant |
| Woźniak et al.15 2021 | PBMC | 0.5, 10, and 100 µM of glyphosate and 250 µM of AMPA | EHMT1, EHMT2 and HDAC | Glyphosate and AMPA | Increase |
| Duforestel et al.9 2019 | MCF10A | 10-11 M (cell concentration used = 2.2×10-6) | TET3 | Glyphosate | Increase |
| Courant et al.16 2022 | MCF10A | 280 µg/L of glyphosate (at least in biological triplicates) | TET3 | Glyphosate | Increase |
Smith et al.14 found that Roundup® decreased the expression of Dnmt1 in the testes of Oryzias latipes at a dose of 5mg/L. No significant variation was observed in the ovaries. In fish larvae, glyphosate was shown to increase the expression of TET1 and TET3 at all the doses tested (0.5 mg/L glyphosate and 0.5 mg/L and 5 mg/L Roundup®), while TET2 expression was increased at 0.5 mg/L and 5 mg/L Roundup®. In addition, the enzyme Dnmt1 had its expression decreased at all doses to which the larvae were exposed.
Larsen et al.5 in their study, pig models exposed to glyphosate at 20 ppm and 200 ppm found no variation in the expression of Dnmt1, Dnmt2, Dnmt3A and Dnmt3B in any of the tissues evaluated (kidney, liver and small intestine) at any dose. In addition, TET3 also showed no significant variation in expression in any of the tissues evaluated. Another study that presented different results to Smith et al.14 is that of Woźniak et al.15. The article used PMBC exposed to glyphosate and AMPA as an experimental model. Both compounds increased the expression of Dnmt1, in the case of the herbicide at doses of 0.5, 10, and 100μM, and with the metabolite at doses of 100 and 250 μM. Dnmt3A, on the other hand, had its expression boosted at the highest concentration of glyphosate. Also, in the work by Woźniak et al.15, they analyzed other enzymes, such as EHMT1 and EHMT2, both of which showed no significant variation in expression, unlike EHMT1, EHMT2 and HDAC which had their expression increased at all doses of glyphosate and at the highest dose of AMPA.
In MCF10A strains, Duforestel et al.9 obtained the following data: the TET3 enzyme had its expression and activity induced by glyphosate at a dose of 10-11 M, results similar to those achieved by Smith et al.14. Along the same lines, Courant et al.16 also found an increase in TET3 expression in MCF10A strains exposed to 280 μg/L of glyphosate.
Hormonal disruption
Glyphosate has been shown to deregulate genes related to hormonal functions in several studies. In Smith et al.14, Gomez et al.17, Lorenz et al.18 and Costa Reis et al.19 it was observed that glyphosate and Roundup® modified methylation and gene expression in different hormonal pathways (Table 3).
Table 3
Comparison of results in relation to hormonal disruption
| Authors Year | Experimental model | Concentration | Compound | Altered gene | Expression |
|---|---|---|---|---|---|
| Smith et al.14 2019 | Oryzias latipes adult females (n=9–12) (brain) | 0.5 mg/L | Roundup® | Gpr54-1 and Gpr54-2 | Decrease |
| Smith et al.14 2019 | Oryzias latipes adult females (n=9–12) (brain) | 5 mg/L | Roundup® | Gpr54-1 | Decrease |
| Smith et al.14 2019 | Oryzias latipes adult males (n=8–16) (brain) | 0.5 mg/L | Roundup® | Fshr and Arα | Decrease |
| Gomez et al.17 2019 | Male Wistar rats (mammary glands) (n=6–8) | 3.7 mg/kg body weight/day | Glyphosate-based herbicide | AR and ESR1 | Decrease |
| Gomez et al.17 2019 | Male Wistar rats (mammary glands) (n=6–8) | 352 mg/kg body weight/day | Glyphosate-based herbicide | ESR1 | Decrease |
| Lorenz et al.18 2018 | Female Wistar rats (uteri) (n=8) | 352 mg/kg body weight/day | Glyphosate-based herbicide | ESR1 | Increase |
| Costa Reis et al.19 2021 | Male Wistar rats* (cerebellum) (n=24) | 50 mg/kg body weight/day | Roundup® | Trα1 | Increase |
| Costa Reis et al.19 2021 | Male Wistar rats (cerebellum) (n=24) | 5 mg/kg body weight/day | Roundup® | Trβ1 | Decrease |
| Costa Reis et al.19 2021 | Male Wistar rats (cerebellum) (n=24) | 5 mg/kg body weight/day | Roundup® | Mct8 | Increase |
| Costa Reis et al.19 2021 | Male Wistar rats (cerebellum) (n=24) | 5 mg/kg body weight/day | Roundup® | Dio3 | Decrease |
Smith et al.14 used female Oryzias latipes as an experimental model. After exposure to the herbicide Roundup®, a decrease in the expression of the kisspeptin receptor genes (Gpr54-1 and Gpr54-2) was observed at a concentration of 0.5 mg/L in brain tissue. However, at a dose of 5mg/L only the Gpr54-1 gene had its expression significantly decreased. Glyphosate at a concentration of 0.5 mg/L did not alter the expression of either of the two genes altered in the presence of Roundup®. The Kiss1 and Kiss2 genes, unlike their receptors, did not undergo any significant change in expression in any of the treatments.
In the same study, Smith et al.14 in Oryzias latipes testicles showed a significant decrease in the expression of the FSH receptor gene (Fshr) and the androgen receptor (Ar ) when exposed to 0.5mg/L of Roundup® and glyphosate. In addition, the Dmrt1 gene showed a drop in expression when treated with glyphosate at a dose of 0.5 mg/L.
As already mentioned, glyphosate is not marketed pure; it is usually glyphosate-based herbicides (HBG). In the work by Gomez et al.17, HBG altered gene expression and methylation status in the mammary glands of male Wistar rats bred and breastfed by females exposed to the herbicide from the ninth day of gestation until weaning. In them, the AR and ESR1 genes (androgen receptor and estrogen receptor, respectively) had a significant decrease in mRNAs at a dose of 3.7 mg/kg body weight/day (3.7 HBG). However, at a higher concentration of 352 mg/kg body weight/day (352 HBG) only ESR1 maintained its significant drop in expression, with AR expression being unchanged. However, even with a decrease in AR mRNAs, AR protein expression increased significantly in the 3.7 HBG group, unlike ESR1 where the presence of its protein also decreased along with the expression of its mRNAs.
The decrease in ESR1 expression was accompanied by a decrease in the expression of specific 5’UTR promoters due to hypermethylation, in line with the decrease in ESR1 transcripts and proteins. The exon called ESR1-OS decreased its expression in rats at the 3.7 HBG dose, explained by the observed increase in methylation status in the same promoter. In rats exposed to 352 HBG, there was a decrease in the expression of the ESR1-O, ESR1-OT and ESR1-E exons, also explained by the observed increase in methylation of the same exons19.
Results referring to another tissue were observed in female Wistar rats in Lorenz et al.18. When ESR1 expression was analyzed in the uteri of females with perinatal exposure to 352 HBG, a significant increase in ESR1 expression was observed in uterine tissues collected during the embryo’s pre-implantation period. In addition, an increase in the transcription of the ESR1-O 5’ UTR exon was observed at the same dose. A more detailed analysis of the ESR1-O promoter showed a decrease in methylation at the BstUI-1 restriction site. Analyzing the post-translational modifications in the histones linked to different restriction sites in the ESR1-O promoter showed a decrease in H3 acetylation at the MaeII site, while histone H4 acetylation increased at all the sites measured (MaeII, BstUI 1 and BstUI 2). In another analysis from the same study, it was observed that H3K9 lysine methylation increased at two sites (MaeII, BstUI). H3K9me3 methylation is linked to strong chromatin condensation20; however, it has a dual role as it is also present in active regions of genes transcribed in mammals21. Finally, H3K27 methylation decreased at all sites evaluated, with H3K27me3 being associated with gene silencing22. Overall, the data suggest that glyphosate deregulates the transcription of the ESR1 gene in female Wistar rats exposed in the perinatal period to a dose of 352 HBG and this effect can possibly be explained by an increase in the expression of the ESR1-O promoter in the 5’UTR region. This increase, in turn, is supposedly explained by an increase in H4 acetylation and a decrease in H3K27 methylation21.
Also analyzed was the hormonal imbalance due to glyphosate in the cerebellum of Wistar rats exposed during the perinatal period to a dose of 50 mg/kg body weight/day of Roundup® (50RU). The animals showed a significant increase in the expression of the thyroid hormone receptor alpha 1 (Trα1) in the organ. Furthermore, when treated at a dose of 5 mg/kg body weight/day (5RU), there was a drop in the expression of the beta thyroid hormone receptor (Trβ1); however, there was no significant change in the expression of beta 2 thyroid hormone (Trβ2). The thyroid hormone transporter (Mct8) also suffered an imbalance when exposed to the agrochemical, in this case, at a dose of 5RU it suffered an increase in transcription; unlike another transporter (Oatp1c1), for which there was no significant change in expression. Finally, in the thyroid hormone converting enzyme (Dio3) there was a decrease in transcription at the 5RU dose. Unlike the other thyroid hormone converting enzymes, there was no statistically significant change in transcription in any treatment.
Tumor-related genes
Glyphosate, Roundup® and AMPA have been shown to affect the expression of genes related to tumors in the studies surveyed in this review (Woźniak et al.7, Woźniak et al.8, Duforestel et al.9, and Almirón et al.23) (Table 4).
Table 4
Comparison of the results obtained in relation to genes linked to tumors
| Authors Year | Experimental model | Concentration | Compound | Gene | Methylation status |
|---|---|---|---|---|---|
| Woźniak et al.7 2019 | PBMC (healthy volunteers, n=4) | 0.5, 10 and 100 μM | Glyphosate | p16 | Unchanged |
| Woźniak et al.7 2019 | PBMC | 100 μM | Glyphosate | p21 | Decrease |
| Woźniak et al.7 2019 | PBMC | 100 μM | Glyphosate | TP53 | Increase |
| Woźniak et al.7 2019 | PBMC | 100 μM | Glyphosate | BCL2 | Unchanged |
| Woźniak et al.7 2019 | PBMC | 0.5, 10 and 100 μM | Glyphosate | CCND1 | Unchanged |
| Woźniak et al.82020 | PBMC | 0.5, 10 and 250 μM | AMPA | p16 | Unchanged |
| Woźniak et al.8 2020 | PBMC | 0.5, 10 and 250 μM | AMPA | p21 | Decrease |
| Woźniak et al.8 2020 | PBMC | 0.5, 10 and 250 μM | AMPA | TP53 | Increase |
| Woźniak et al.8 2020 | PBMC | 0.5, 10 and 250 μM | AMPA | BCL2 | Unchanged |
| Woźniak et al.8 2020 | PBMC | 0.5, 10 and 250 μM | AMPA | TP53 | Unchanged |
| Duforestel et al.9 2019 | MCF10A | 10-11 M (cell concentration used = 2.2×10-6) | Glyphosate | DUX4 and MTRNR2L2 | Decrease |
| Almirón et al.23 2024 | Female Wistar rats* (uterus) | 3.8 and 3.9 mg/kg body weight/day, respectively | Glyphosate and Magnum Super III | Lif | Increase |
In Woźniak et al.7, exposure of PMBC to glyphosate altered the expression of tumor suppressor genes. At doses of 0.5, 10 and 100 μM of glyphosate there was a significant drop in the expression of the p16 protein; however, this drop was not accompanied by a significant change in the percentage of methylation of the gene. At the highest dose (100 μM) there was an increase in p21 transcription and a decrease in TP53 expression. In relation to the methylation status of the same genes, it was observed that the p21 gene showed a decrease in methylation at all treated concentrations, while the TP53 gene showed an increase in methylation at all concentrations. Proto-oncogenes are also affected by exposure to glyphosate in PMBC. The BCL2 proto-oncogene had its expression increased in cells exposed to a dose of 100 μM, even though the percentage of methylation did not change. The CCND1 gene also had its expression increased at all the doses tested, even though the percentage of methylation was not significantly altered at any dose. It can be seen that the change was not accompanied in all cases by a change in the methylation of the gene. This was observed in the p16, BCL2 and CCND1 genes. Another point to note is that Woźniak et al.7 showed a drop in the percentage of global 5mC, even though the genes analyzed did not have altered methylation.
Woźniak et al.8 also found that the metabolite AMPA altered the expression and methylation of tumor suppressor genes and proto-oncogenes in PMBC. In this study, p16 showed no change in expression or methylation status at any of the doses tested (0.5, 10 and 250 μM). In p21, the methylation status decreased at all the concentrations tested, but there was no change in the gene’s expression. Finally, TP53 had its methylation increased at all the doses tested, but there was still no change in expression. In relation to proto-oncogenes, the BCL2 gene had no altered expression or methylation at any concentration, while CCND1 had its expression increased in all treatments; however, this increase was not accompanied by a change in methylation.
Figure 1
Eligibility criteria for the selection of articles identified in the databases and used in the PRISMA-based study4
In the work by Duforestel et al.9 two genes were found to be hypomethylated after exposure to glyphosate in MCF10A cells at a dose of 10-11M, namely DUX4 and MTRNR2L2. These are also shown to interact with the TET3 enzyme, which, as already mentioned, is responsible for the hypomethylation process, by converting 5mC into 5hmC (5 hydroxy-methylcytosine), which is subsequently transformed into a demethylated cytosine; these mechanisms of epigenetic modification of the TETs (TET1, TET2 and TET3) are linked to tumorigenesis24.
In another analysis by Duforestel et al.9, MCF10A cells were treated with glyphosate and a microRna (miR182-5p), a mimetic single-stranded regulatory microRna that is deregulated in a number of human tumors. After treatment, the cells were injected subcutaneously into four Wistar rats. Of these four, two developed tumors. Interestingly, the cells treated only with glyphosate or with miR182-5p did not develop tumors.
In order to analyze the relationship between the TET3 enzyme and the genetic imbalance of DUX4 and MTRNR2L2 and relate it to tumor formation, another analysis was carried out. The authors treated MCF10A cells with glyphosate and miR182-5p and exposed them to DMOG at a dose of 1 mM, a compound that inhibits TET3 activity. As a result, no tumors appeared in the mice that received the cells.
Almirón et al.23 found that female Wistar rats exposed during the perinatal period had a drop in the expression of the Lif leukemia inhibitor gene in uterine tissues. The drop was seen in treatments with glyphosate and with a glyphosate-based herbicide (Magnum super III®) at a dose of 3.8 and 3.9 mg /kg body weight/day, respectively. A broader analysis of the Lif gene and its regulatory regions (RR1, RR2 and RR3) all showed an increase in methylation, both when exposed to glyphosate and glyphosate-based herbicide. The relationship between these regulatory regions and histones shows that in H3 there was a significant increase in acetylation in RR1 and RR3 in groups exposed to glyphosate-based herbicide. In the group exposed to glyphosate, there was a decrease in H3 acetylation in all RRs. There was also an increase in H3K27 trimethylation in the RR1 and RR3 regulatory regions when exposed to glyphosate.
From the results of the studies, it can be concluded that glyphosate deregulated the gene expression or methylation status of genes linked to tumors, generally contributing to the silencing of tumor suppressor genes. It is also observed that glyphosate influences the functioning of the TET3 enzyme, which in turn is linked to the hypomethylation of DUX4 and MTRNR2L2, contributing to the activation of proto-oncogenes, and it is parsimonious to assume that it may be linked to hypomethylation in more genes.
Action of glyphosate on the Hoxa10 gene
Both glyphosate and glyphosate-based herbicide (HBG) were shown to alter Hoxa10 gene expression in the work by Lorenz et al.25. In the article, the gene’s expression was reduced in the uterine tissues of Wistar rats exposed during the perinatal period to 3.8 HBG mg/kg body weight/day and 3.9 glyphosate mg/kg body weight/day. The same study also provided a more in-depth analysis of the gene’s methylation status. In all treatments, the gene was hypermethylated in the four regulatory regions (RR) evaluated. The post-translational modifications of the histones linked to the RRs were also evaluated. HBG increased the acetylation of H3, H4 (in H4 RR2 there was a decrease) and the methylation of H3K27 in RR1, RR3 and RR4. Glyphosate increased H4 acetylation and H3K27 methylation. However, when Lorenz et al.25 evaluated the expression of enzymes linked to epigenetic mechanisms, many did not show altered expression, such as Dnmt1, Dnmt3A, Dnmat3B, Hdac1 and Hdac3. The only enzyme evaluated that had a change in expression was EZH2, a histone methyltransferase. The enzyme showed an increase in expression in all treatments, somewhat corroborating the increase in H3K27 methylation in all treatments.
In another study, Guerreiro et al.26 also analyzed the Hoxa10 gene. In the article, rats were exposed after birth for seven days via neck injections at a dose of 2 mg/kg body weight/day of Roundup®. In the article, the expression of Hoxa10 decreased after treatment with Roundup®. Interestingly, corroborating the findings of Lorenz et al.25. Another analysis was the RR methylation status of the gene, with an increase in methylation being observed in 4 (S1A, S2, S3 and S4) of the 5 regions that were analyzed. Post-translational modifications of the histones linked to the RRs were also evaluated, with an increase in H3K9me3 methylation in S1A, S1B, S3 and S4 PMBCa decrease in H3K27me3 in all RRs.
Differentiated gene expression
Articles by Jia et al.27, Lian et al.28 and Babich et al.29 showed that glyphosate altered the expression of several classes of genes. Jia et al.27 showed that exposure to glyphosate in Oreochromis niloticus (Tilapia) at a dose of 2 mg/L altered gene expression in livers. After exposure, around 94 genes had their expression increased and 131 had their expression decreased. Among them, the main types of genes altered were metabolic processes, biological regulation, response to stimuli, transport, catalytic activity and binding proteins. All of them had at least 20 related genes altered. In another analysis by Jia et al.27, one of the main metabolic pathways altered by exposure to glyphosate was the PPAR signaling pathway, a pathway linked to lipid metabolism. Among the genes in the pathway that had their expression decreased were PPARα, fads2, cyp8b1, fabp3, acsl4 and angptl4. Only one gene in the pathway had its expression increased, being cpt1a. It is therefore possible that glyphosate alters the normal metabolism of lipids, thus potentially causing health problems. Lian et al.28 showed that low doses of glyphosate increase the risk of hepatic steatosis, while high doses cause liver damage and disrupt lipid metabolism, contributing to the appearance of hepatic steatosis in roosters. An analysis of gene expression showed an increase in the expression of genes linked to the KEGG autophagy pathway, while groups exposed to glyphosate also had a decrease in the expression of genes linked to lipid metabolism, such as PPARα, cpt1a, FABP1 and CD36.
In another study, Babich et al.29 showed that glyphosate also altered the expression of several genes in another experimental model, in this case zebrafish. Fish treated at a dose of 10 ppb had 46 genes differentially expressed. The main classes of genes affected were genes related to DNA remodeling, organization of the extracellular matrix and genes linked to lysosomal processes.
Summary
Global methylation
The data found in the articles show that glyphosate and its metabolite AMPA modify the overall methylation pattern. It is not possible to define a general response to exposure to the agrochemical, given that factors such as route of exposure, dose, experimental model, and tissue analyzed will alter the results found. However, the data collected allow us to understand that in more sensitive models, such as cultured cells and fluids (in this case, from roosters), glyphosate and AMPA cause global hypomethylation. Thus, in the study by Duforestel et al.9, even using a dose lower than the NOAEL (10-11 M), the cells responded significantly to the doses of glyphosate and AMPA30-32. The non-significance in the data obtained with pigs in the study by Larsen et al.5 may be linked to the low dose administered or to the animal’s own organism. It can be observed that, among the articles surveyed in this review, there is no pattern in relation to dose, experimental model, route of exposure, and tissue analyzed. Thus, comparing the data becomes more difficult.
Differentially methylated regions
With the transgenerational analysis by Kubsad et al.11, it can be stated that even with the F0 generation being exposed to glyphosate, we cannot say that the methylation pattern will be the same in other generations. Even with these data, it cannot be confirmed that the epigenetic modifications that occurred in past generations cannot be passed on to future generations. The other studies in the present review show that an F0 generation of rats treated with glyphosate generates an F1 generation, which subsequently generates an F3 generation, which have specific DMR patterns for different diseases. The data obtained show a low overlap between the DMRs of the F3 controls and groups with specific F3 diseases, thus showing that DMRs are likely to be directly linked to diseases.
Enzymes related to epigenetic mechanisms
The articles obtained show that Roundup®, glyphosate, and AMPA alter the expression pattern of genes linked to epigenetic mechanisms. Dnmt1, Dnmt2, and Dnmt3A had increased expression in PMBC; however, Dnmt1 had decreased expression with Roundup®. In addition, in pigs, there was also no significance in the data for these enzymes. These differences, in the first case, can be explained by the fact that Roundup® is a commercial solution with other compounds in its formulation, or by the experimental model, given that the model used in the work of Smith et al.14 was the fish Oryzias latipes. Therefore, for a better understanding of the difference in expression between glyphosate and Roundup® for the same gene, further analysis will be necessary. In the case of pigs, in the work of Larsen et al.15, the experimental model used may be the reason for the difference between the data. Thus, again, the lack of standardization in the model used makes it difficult to compare the data.
The increase in the expression of DNA methyltransferases presented in the work of Smith et al.14 is inconsistent with the data obtained on global methylation, given that, in general, global methylation declined. The reason for the decrease in global methylation may therefore be linked to the TET1, TET2, and TET3 enzymes. Their expression increased in treatments with glyphosate and Roundup®. It is therefore possible to assume that the reason for the global hypomethylation observed in other articles may be linked to TETs, even if the experimental models are not the same.
Hormonal disruption
Data collected on hormonal disruption show that in female Oryzias latipes, Roundup® and glyphosate decrease the expression of Kisspeptin Gpr54-1 and Gpr54-2 receptors in the brain. However, glyphosate alone only had an effect at higher doses, and only on the Gpr54-1 gene. This difference can be explained by the fact that Roundup® does not have a pure formulation, so other compounds may be acting to add to this observed result, which is, once again, a limitation found in the use of formulated glyphosate. In addition to Kisspeptin receptors, FSH receptors, androgen hormone, and the transcription factor Dmtr1 also showed a decrease in expression with exposure to Roundup® and glyphosate, but now in the testes of Oryzias latipes.
Kisspeptin is linked to reproduction, modulating GnRH (gonadotropin-releasing hormone) secretion to play a significant role in the HPG (hypothalamic-pituitary-gonadal) axis, which in turn regulates follicular development, egg maturation, ovulation, and spermatogenesis. Furthermore, with the decrease in Dmtr1 expression, it is possible to correlate the effect of the herbicide on spermatogenesis and sexual differentiation, since the Dmtr1 gene is linked to this process.
In rats, it is possible to correlate exposure to glyphosate-based herbicides with hormonal imbalance, but the imbalance observed in the extracted data shows that the expression profile is not the same between males and females and between different tissues. The decrease in ESR1 and Ar expression occurred in the mammary glands of males that were exposed during gestation. However, in another study obtained in the review, ESR1 expression increased in female uteri. This increase suggests, along with other data collected in the review, that this increase in uteri results from an increase in H4 acetylation at all restriction sites in the exons studied, in addition to decreased methylation. This suggests that the mechanism of action of glyphosate (or glyphosate-based formulations) in the body is diverse and will vary between sexes and tissues.
The data relating to the Trα1, Trβ1, Trβ2, Mct8, Oatp1c1, and Dio3 genes are inconclusive. The data obtained in the review show that there was no pattern of altered expression. In the cerebellums of Wistar rats, Trα1 expression increased when exposed to 50RU, while Trβ1 expression decreased at 5RU. Trβ2 expression was not altered. Each of the Mct8, Oatp1c1, and Dio3 genes had a distinct expression pattern. However, even without a pattern, exposure caused an imbalance in the expression of the genes mentioned above. Thus, as the Trα1 gene is related to neurological development, acting by controlling Purkinje and Bergman cells, specific neurons, and glial cells, the altered expression of Trα1 may possibly lead to failures in cerebellar development33,34.
Tumor-related genes
The data evaluated showed that glyphosate decreased the expression of tumor suppressor genes in PMBCs, such as p16 and TP53, and increased the expression of proto-oncogenes, such as BCL2 and CCND1. However, methylation status did not always change, as in the case of proto-oncogenes and p16. Even though overall methylation decreased, suggesting that post-translational steps in histones may be linked to changes in expression patterns. Furthermore, the metabolite AMPA also caused an increase in CCND1 expression, but without any change in methylation.
The work of Duforestel et al.9 managed to show a complex interaction between glyphosate and other tumorigenic factors, demonstrating that glyphosate can act synergistically with other molecules, such as the aforementioned microRNA (miR182-5p), in the appearance of a tumor. This suggests that glyphosate may not be the only factor triggering a tumor, but it is an important agent in this process. Furthermore, the study showed a relationship between the TET3 enzyme and the appearance of tumors in scenarios of exposure to glyphosate and miR182-5p, such that the inhibition caused by DMOG prevented the generation of tumors in rats. This corroborates the findings of the topics ‘Global methylation’ and ‘Enzymes related to epigenetic mechanisms’, where enzymes of the TET family had their expression increased with exposure to glyphosate and Roundup®, and these enzymes may be linked to global hypomethylation.
Finally, the results obtained by Almirón et al.23 show that glyphosate and HBG may be linked to leukemia. Exposure to the agrochemical resulted in increased methylation in regulatory regions. However, there was a difference in the results linked to H3 acetylation with HBG and glyphosate. This suggests, once again, that the degree of purity of the compounds influences the results obtained.
Action of glyphosate on the Hoxa10 gene
The articles obtained showed that glyphosate, glyphosate-based herbicides, and Roundup® are capable of altering the expression and methylation of Hoxa10. In this case, decreasing gene expression and increasing methylation in the regulatory regions of the gene. Hoxa10 is an important reproductive gene, and a decrease in reproductive capacity and disorders in sexual differentiation can be assumed. However, more experimental data are needed to confirm the physiological results of this imbalance.
Differentiated gene expression
The articles included in the review provided data showing that some genes of interest to the scope of the review had increased expression, as in the case of genes responsible for telomere organization and maintenance, genes linked to negative gene regulation, gene silencing, and chromatin silencing. This shows the relationship between glyphosate and a disruption of epigenetic mechanisms, as has already been demonstrated in other articles cited during the review.
Limitations
In addition to the lack of standardization in doses and experimental models reported in the original sources, we note a recurring difficulty in distinguishing the effects of pure glyphosate from those of commercial glyphosate-based herbicides (HBG) like Roundup®. Moreover, the mechanism of action and routes of administration for glyphosate were highly diverse and vary significantly among the included studies, which makes it difficult to establish a single, overarching pattern of epigenetic response. Finally, as the review was limited to articles published in English between 2018 and 2024, we may have excluded relevant earlier research or studies published in other languages. Additionally, some studies were excluded because they did not analyze glyphosate independently, making it impossible to isolate its specific toxicogenomic effects.
CONCLUSION
Our review noted that glyphosate, in general, may lead to a decrease in global methylation, variation in DMRs in important gene classes such as metabolism, decreased gene expression of hormone receptors, increased expression of the enzyme TET3 and DNMT1 and imbalance in lipid metabolism; however with broad variability noted across studies. In addition, epigenetic mechanisms were noted to be related to altered expression in cancer-related genes following exposure to glyphosate. These results were observed at different doses and targets; however, the interaction of glyphosate with each organism is complex. Further studies are needed to elucidate the epigenetic and toxicogenomic effects of glyphosate.


